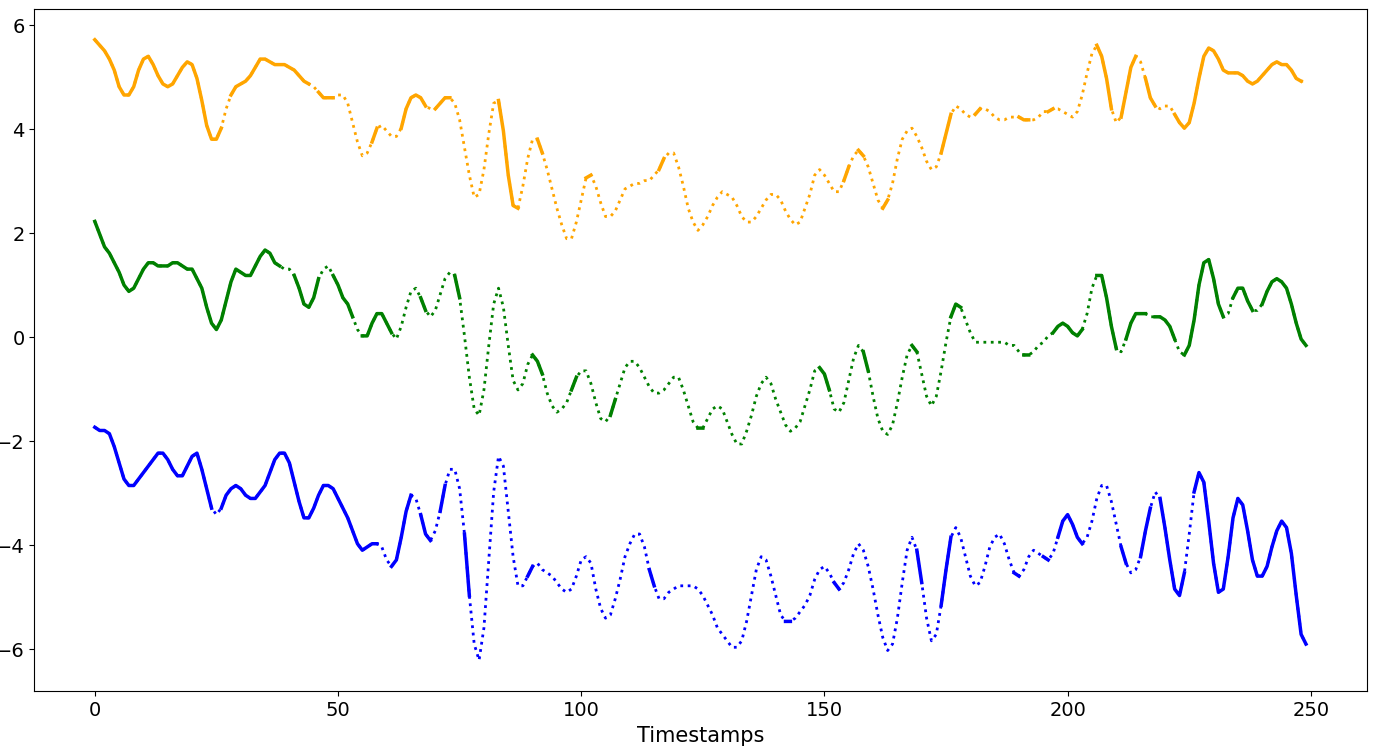
Missingness Patterns¶
ImputeGAP introduces a new taxonomy of missingness patterns tailored to time series, going beyond the traditional MAR and MNAR categories, which were not designed for temporal data.
Setup¶
Note
(N, M) : number of timestamps, number of series
W : user-defined offset window in the beginning of the series (default = 25)
R : user-defined % of missing values (default = 20%)
S : user-defined % of contaminated series (default = 20%)
MONO-BLOCK¶
One missing block per series
Aligned
Missing blocks start at the same selected positions and have the same fixed size across the chosen series, resulting in aligned missing intervals.
Note
R ∈ [1%, (100-W)%]The size of a single missing block varies between 1% and (100 -
W)% ofN.The starting position is the same and begins at
Wand progresses until the size of the missing block is reached, affecting the first series from the top up toS%of the dataset.GenGap.aligned(ts.data, rate_dataset=1, rate_series=0.4, offset=25)
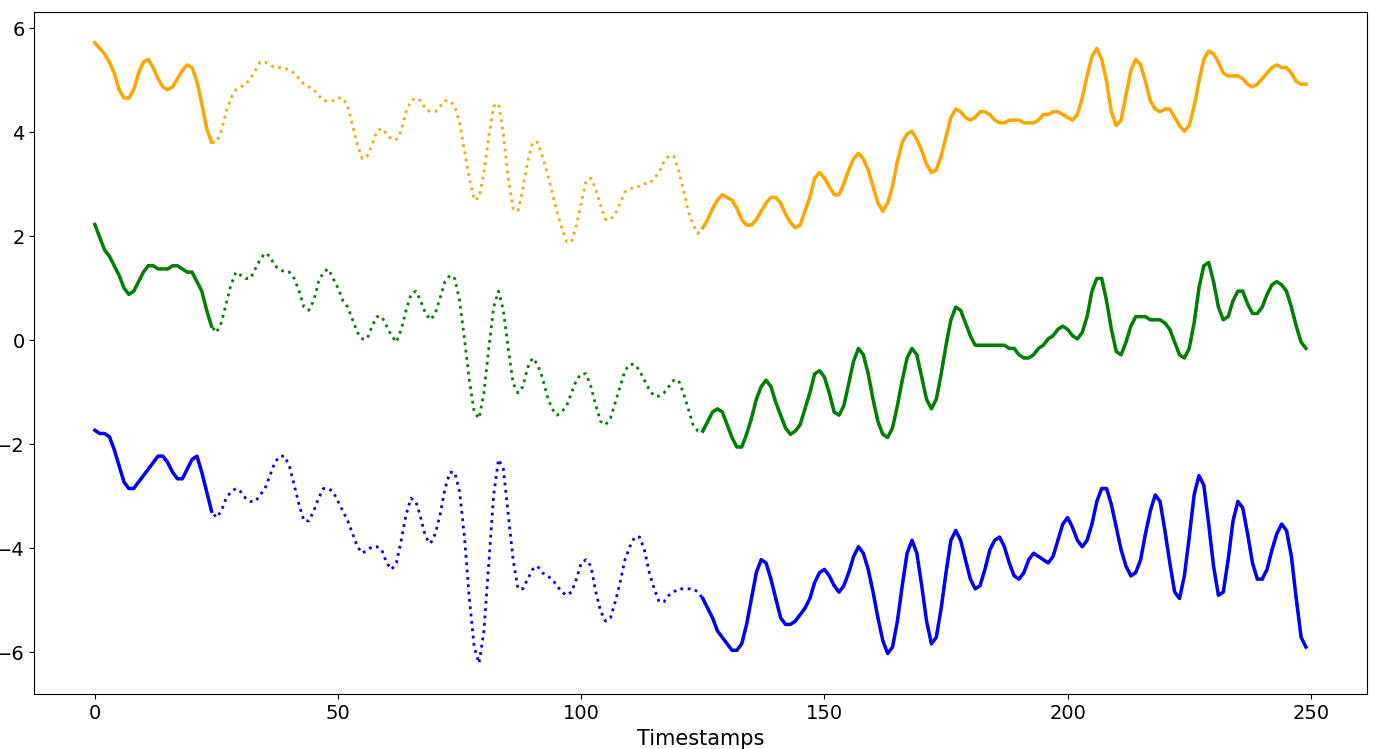
Disjoint
Each missing block begins where the previous one ends, so the missing intervals are consecutive and do not overlap.
Note
R ∈ [1%, (100-W)%]The size of a single missing block varies between 1% and (100 -
W)% ofN.The starting position of the first missing block begins at
W.GenGap.disjoint(ts.data, rate_series=0.4, offset=25)
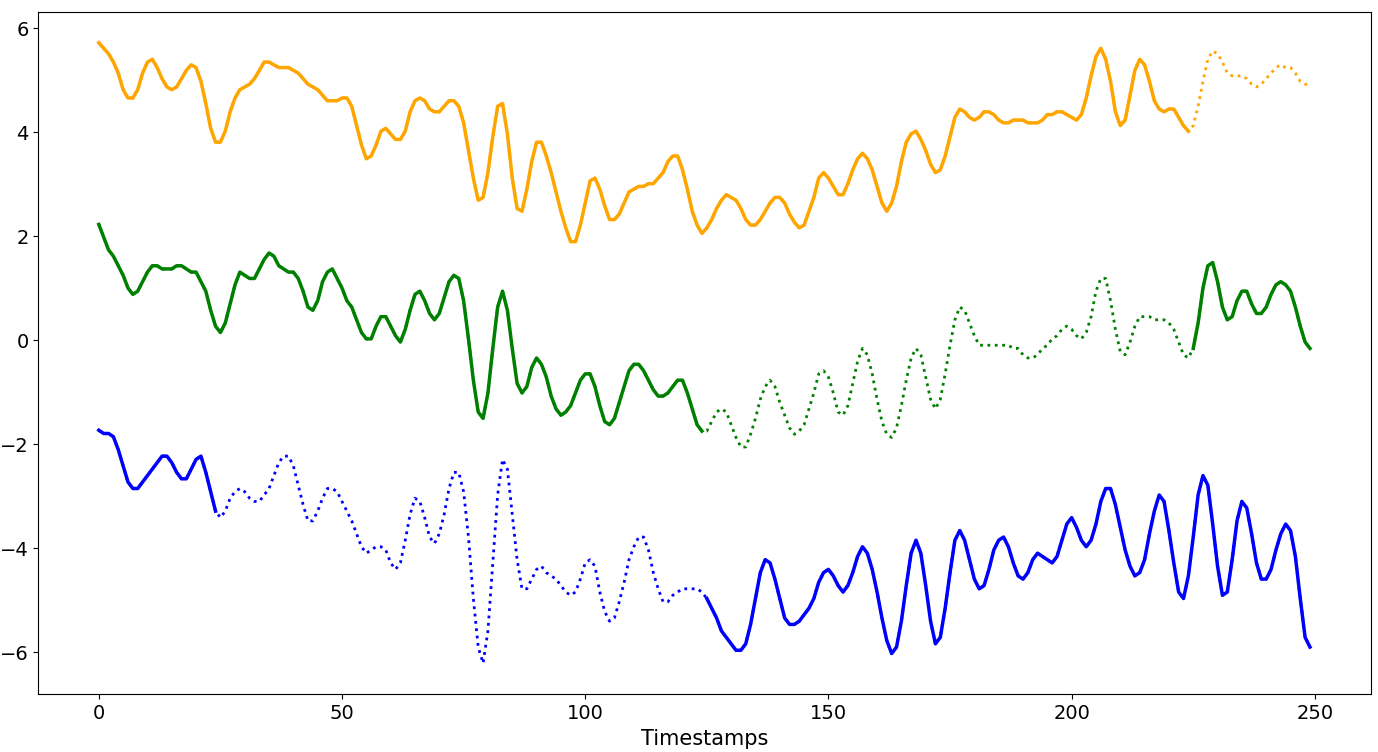
Overlap
Each missing block overlaps with the previous one.
Note
R ∈ [1%, (100-W)%]The size of a single missing block varies between 1% and (100 -
W)% ofN.The starting position of the first missing block begins at
W.The overlap is controlled by the variable
shift.This pattern continues until the limit or
Nis reached.GenGap.overlap(ts.data, rate_series=0.4, offset=25, shift=0.1)
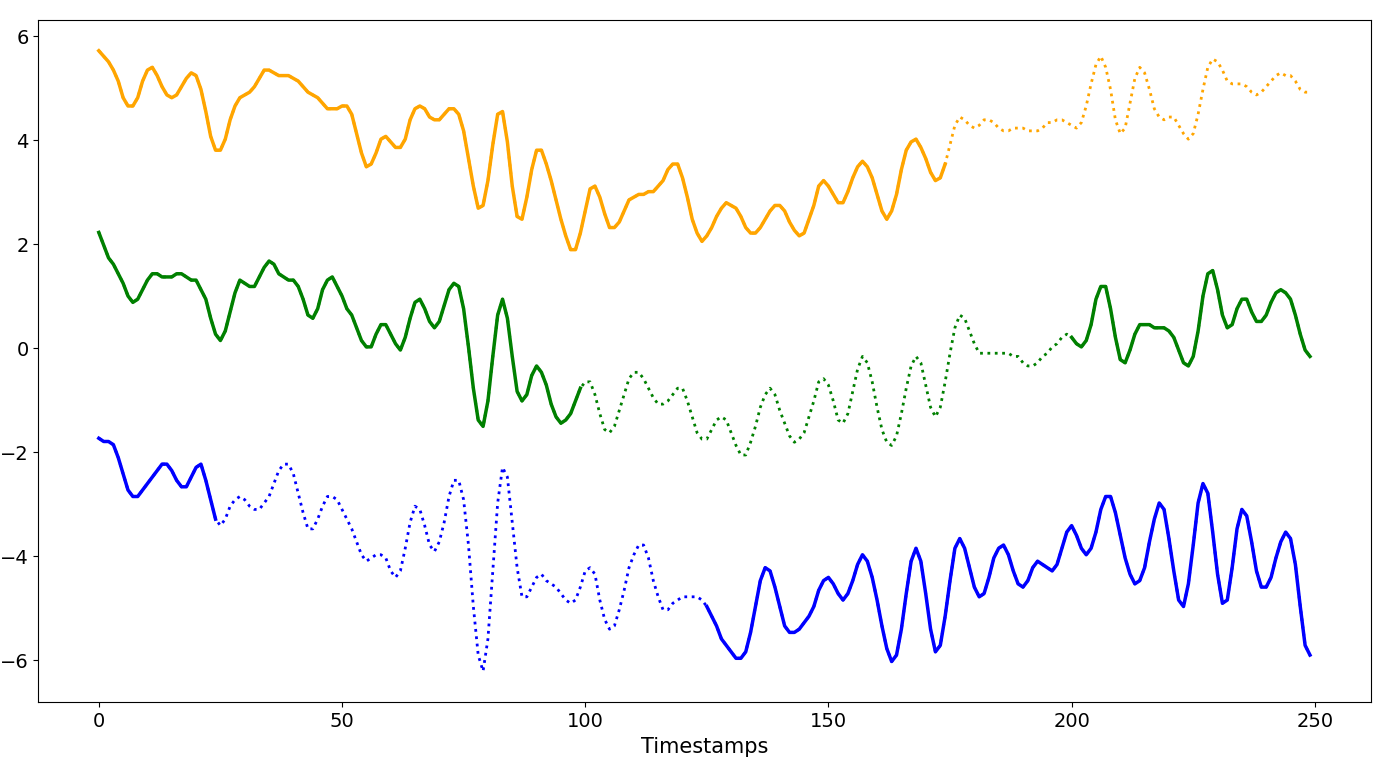
Scattered
The starting position of the missing block is chosen at random, all missing blocks share the same size.
Note
R ∈ [1%, (100-W)%]The size of a single missing block varies between 1% and (100 -
W)% ofN.The starting position is random, then progresses until the size of the missing block is reached, affecting the first series from the top up to
S%of the dataset.GenGap.scattered(ts.data, rate_dataset=1, rate_series=0.4, offset=25)
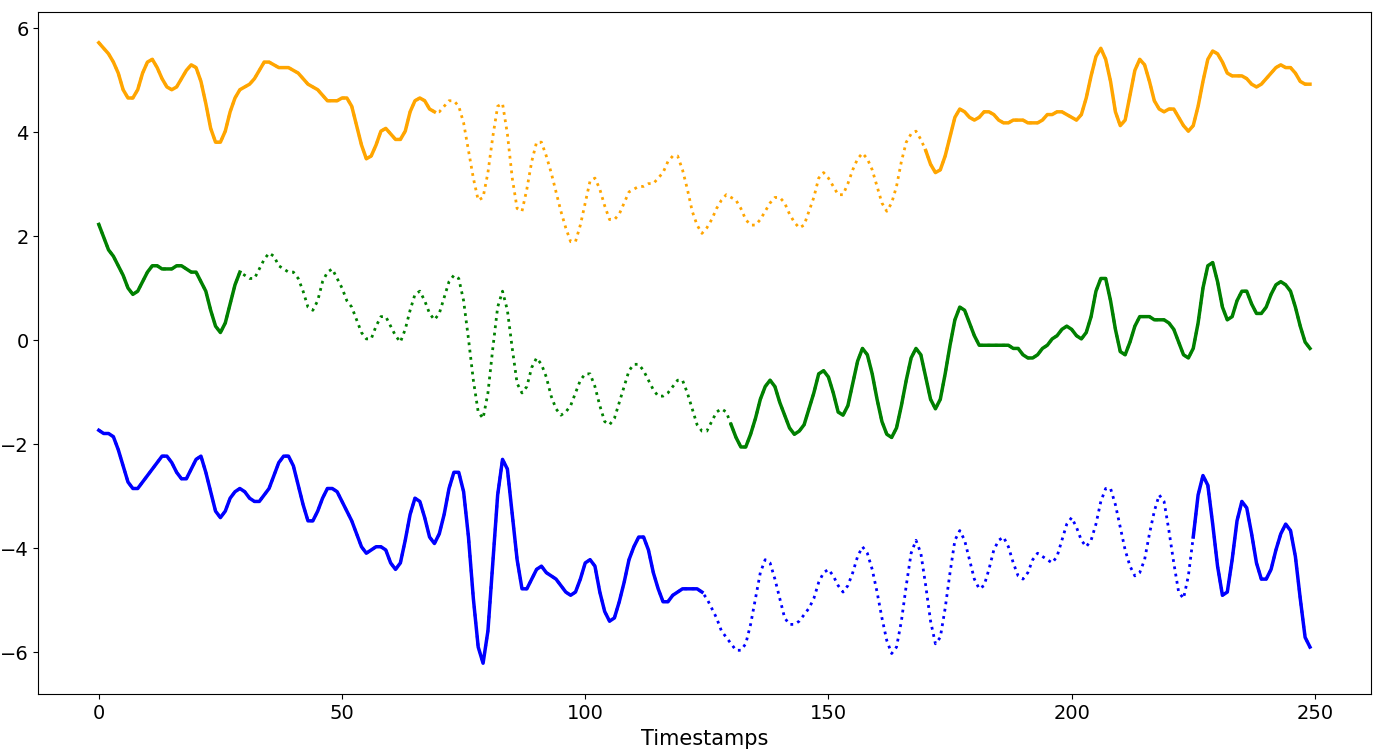
MULTI-BLOCK¶
Multiple missing blocks per series
MCAR
Missing blocks have the same size and are introduced completely at random. The affected time series are selected at random.
Note
R ∈ [1%, (100-W)%]Data blocks of the same size are removed from arbitrary series at a random position between
WandN, until the total number of missing values per series is reached.GenGap.mcar(ts.data, rate_dataset=1, rate_series=0.2, offset=25, seed=False, block_size=20)
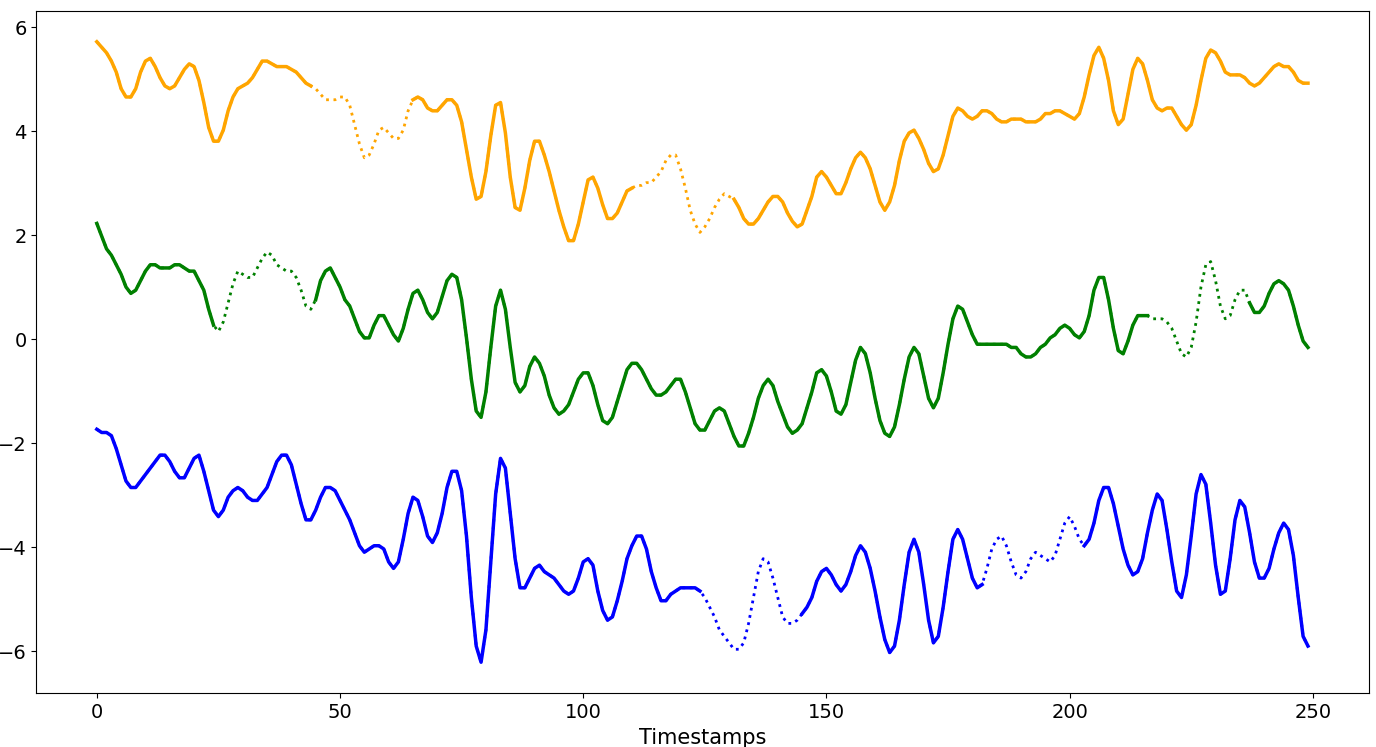
Block Distribution
Missing data follows a probability distribution, each position has a certain chance of being missing.
Note
R ∈ [1%, (100-W)%]Data is removed following a distribution given by the user for every values of the series, affecting the first series from the top up to
S%of the dataset.GenGap.gaussian(ts.data, rate_dataset=1, rate_series=0.4, offset=25, selected_mean="position", std_dev=0.2)
To configure the block distribution pattern, please refer to this page.